import sys
import os
sys.path.append(os.path.abspath('..'))
import libpysal
libpysal.examples.available()
libpysal.examples.explain('mexico')
import geopandas
pth = libpysal.examples.get_path("mexicojoin.shp")
gdf = geopandas.read_file(pth)
from libpysal.weights import Queen, Rook, KNN
%matplotlib inline
import matplotlib.pyplot as plt
ax = gdf.plot()
ax.set_axis_off()
w_rook = Rook.from_dataframe(gdf)
w_rook.n
w_rook.pct_nonzero
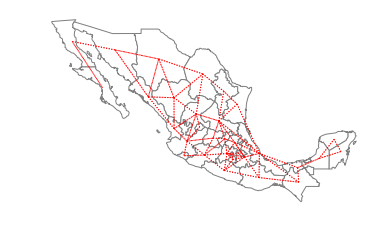
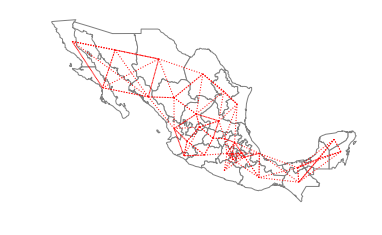
ax = gdf.plot(edgecolor='grey', facecolor='w')
f,ax = w_rook.plot(gdf, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
gdf.head()
w_rook.neighbors[0] # the first location has two neighbors at locations 1 and 22
gdf['NAME'][[0, 1,22]]
So, Baja California Norte has 2 rook neighbors: Baja California Sur and Sonora.
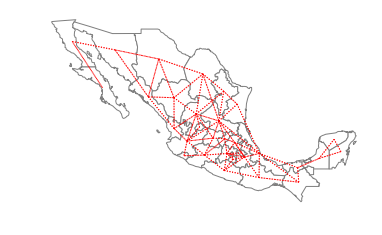
Queen neighbors are based on a more inclusive condition that requires only a shared vertex between two states:
w_queen = Queen.from_dataframe(gdf)
w_queen.n == w_rook.n
(w_queen.pct_nonzero > w_rook.pct_nonzero) == (w_queen.n == w_rook.n)
ax = gdf.plot(edgecolor='grey', facecolor='w')
f,ax = w_queen.plot(gdf, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
w_queen.histogram
w_rook.histogram
c9 = [idx for idx,c in w_queen.cardinalities.items() if c==9]
gdf['NAME'][c9]
w_rook.neighbors[28]
w_queen.neighbors[28]
import numpy as np
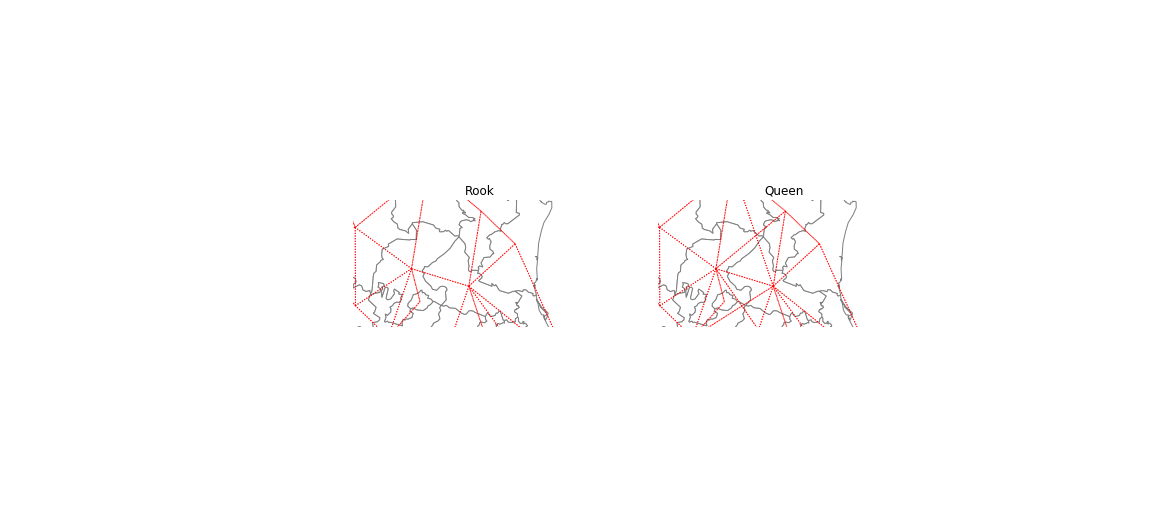
f,ax = plt.subplots(1,2,figsize=(10, 6), subplot_kw=dict(aspect='equal'))
gdf.plot(edgecolor='grey', facecolor='w', ax=ax[0])
w_rook.plot(gdf, ax=ax[0],
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax[0].set_title('Rook')
ax[0].axis(np.asarray([-105.0, -95.0, 21, 26]))
ax[0].axis('off')
gdf.plot(edgecolor='grey', facecolor='w', ax=ax[1])
w_queen.plot(gdf, ax=ax[1],
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax[1].set_title('Queen')
ax[1].axis('off')
ax[1].axis(np.asarray([-105.0, -95.0, 21, 26]))
w_knn = KNN.from_dataframe(gdf, k=4)
w_knn.histogram
ax = gdf.plot(edgecolor='grey', facecolor='w')
f,ax = w_knn.plot(gdf, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
pth = libpysal.examples.get_path("mexicojoin.shp")
from libpysal.weights import Queen, Rook, KNN
w_queen = Queen.from_shapefile(pth)
w_rook = Rook.from_shapefile(pth)
w_knn1 = KNN.from_shapefile(pth)
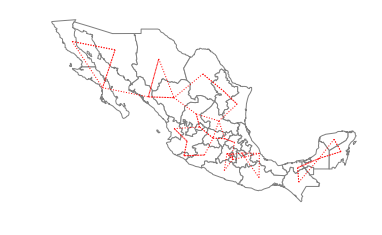
The warning alerts us to the fact that using a first nearest neighbor criterion to define the neighbors results in a connectivity graph that has more than a single component. In this particular case there are 2 components which can be seen in the following plot:
ax = gdf.plot(edgecolor='grey', facecolor='w')
f,ax = w_knn1.plot(gdf, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
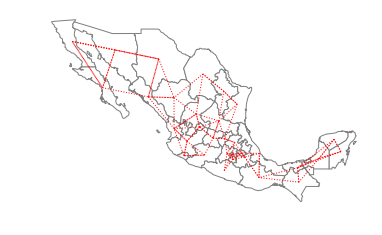
The two components are separated in the southern part of the country, with the smaller component to the east and the larger component running through the rest of the country to the west. For certain types of spatial analytical methods, it is necessary to have a adjacency structure that consists of a single component. To ensure this for the case of Mexican states, we can increase the number of nearest neighbors to three:
w_knn3 = KNN.from_shapefile(pth,k=3)
ax = gdf.plot(edgecolor='grey', facecolor='w')
f,ax = w_knn3.plot(gdf, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
from libpysal.weights import lat2W
w = lat2W(4,3)
w.n
w.pct_nonzero
w.neighbors
rs = libpysal.examples.get_path('map_RS_BR.shp')
import geopandas as gpd
rs_df = gpd.read_file(rs)
wq = libpysal.weights.Queen.from_dataframe(rs_df)
len(wq.islands)
wq[0]
wf = libpysal.weights.fuzzy_contiguity(rs_df)
wf.islands
wf[0]
plt.rcParams["figure.figsize"] = (20,15)
ax = rs_df.plot(edgecolor='grey', facecolor='w')
f,ax = wq.plot(rs_df, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_axis_off()
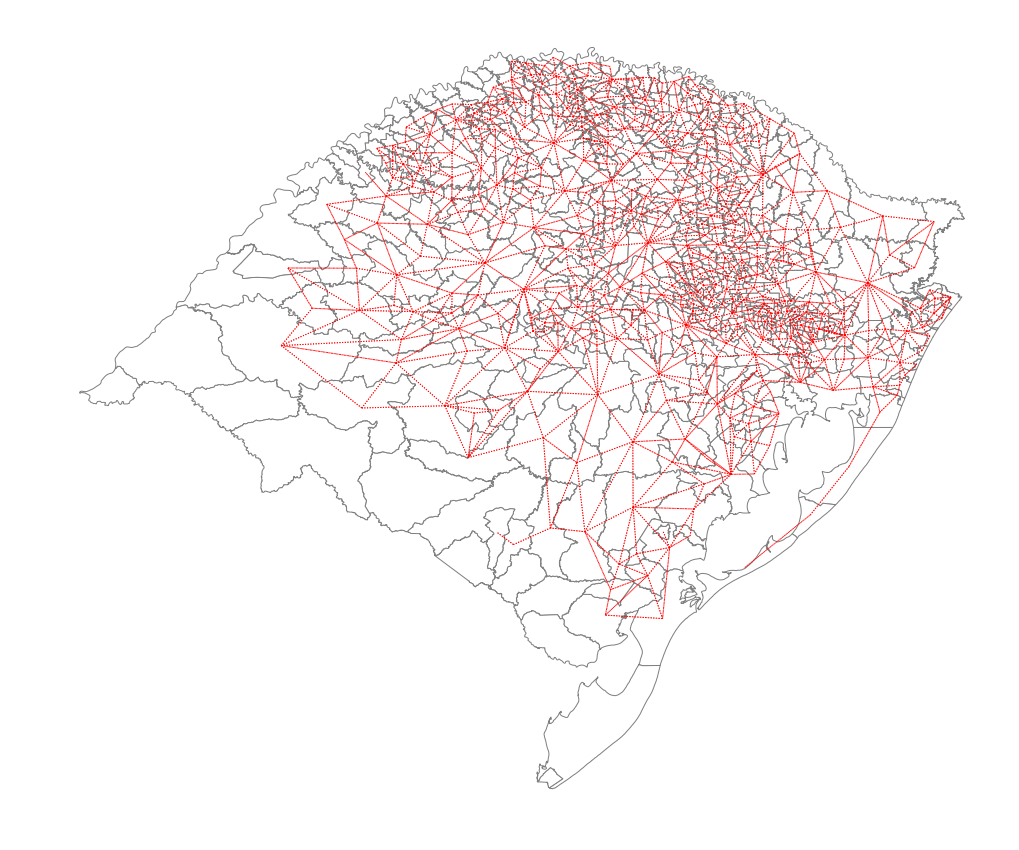
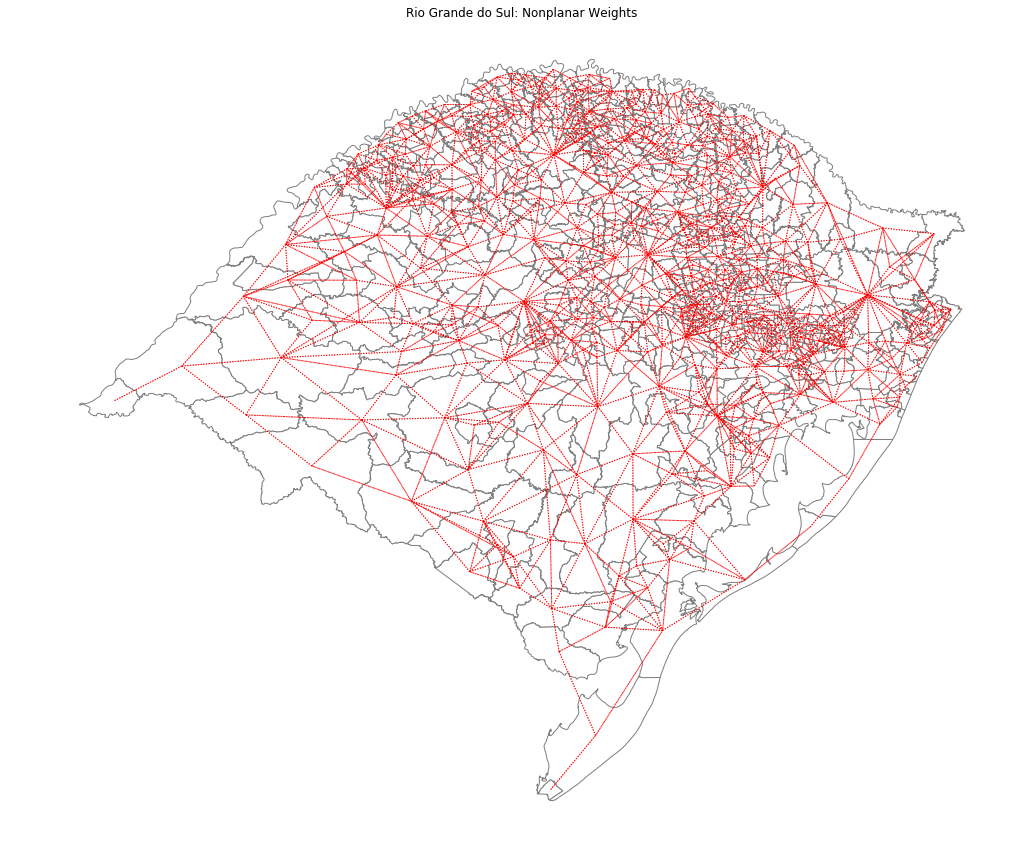
ax = rs_df.plot(edgecolor='grey', facecolor='w')
f,ax = wf.plot(rs_df, ax=ax,
edge_kws=dict(color='r', linestyle=':', linewidth=1),
node_kws=dict(marker=''))
ax.set_title('Rio Grande do Sul: Nonplanar Weights')
ax.set_axis_off()
plt.savefig('rioGrandeDoSul.png')