Voronoi Polygons for 2-D Point Sets¶
Author: Serge Rey (http://github.com/sjsrey)
Basic Usage¶
import os
import sys
sys.path.append(os.path.abspath(".."))
from libpysal.cg.voronoi import voronoi, voronoi_frames
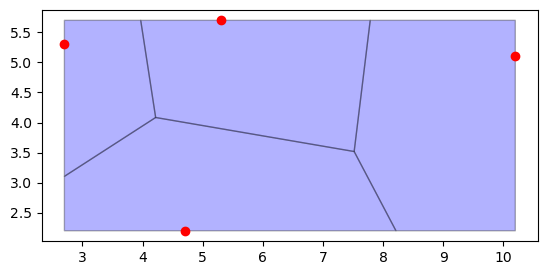
points = [(10.2, 5.1), (4.7, 2.2), (5.3, 5.7), (2.7, 5.3)]
regions, vertices = voronoi(points)
/tmp/ipykernel_4300/2901581476.py:1: FutureWarning: The 'voronoi' function is considered private and will be removed in a future release.
regions, vertices = voronoi(points)
/home/runner/micromamba/envs/test/lib/python3.14/site-packages/libpysal/cg/voronoi.py:67: FutureWarning: The 'voronoi_regions' function is considered private and will be removed in a future release.
vor = voronoi_regions(Voronoi(points), radius=radius)
regions
[[1, 3, 2], [4, 5, 1, 0], [0, 1, 7, 6], [9, 0, 8]]
vertices
array([[ 4.21783296, 4.08408578],
[ 7.51956025, 3.51807539],
[ 9.4642193 , 19.3994576 ],
[ 14.98210684, -10.63503022],
[ -9.22691341, -4.58994414],
[ 14.98210684, -10.63503022],
[ 1.78491801, 19.89803294],
[ 9.4642193 , 19.3994576 ],
[ 1.78491801, 19.89803294],
[ -9.22691341, -4.58994414]])
region_df, point_df = voronoi_frames(points)
/tmp/ipykernel_4300/2172163060.py:1: FutureWarning: The 'as_gdf' parameter currently defaults to True but will default to False in a future release. Set it explicitly to avoid this warning.
region_df, point_df = voronoi_frames(points)
/tmp/ipykernel_4300/2172163060.py:1: FutureWarning: The 'return_input' parameter currently defaults to True but will default to False in a future release. Set it explicitly to avoid this warning.
region_df, point_df = voronoi_frames(points)
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
fig, ax = plt.subplots()
region_df.plot(ax=ax, color="blue", edgecolor="black", alpha=0.3)
point_df.plot(ax=ax, color="red")
<Axes: >
Larger Problem¶
n_points = 200
np.random.seed(12345)
points = np.random.random((n_points, 2)) * 10 + 10
results = voronoi(points)
mins = points.min(axis=0)
maxs = points.max(axis=0)
/tmp/ipykernel_4300/3060736572.py:4: FutureWarning: The 'voronoi' function is considered private and will be removed in a future release.
results = voronoi(points)
/home/runner/micromamba/envs/test/lib/python3.14/site-packages/libpysal/cg/voronoi.py:67: FutureWarning: The 'voronoi_regions' function is considered private and will be removed in a future release.
vor = voronoi_regions(Voronoi(points), radius=radius)
regions, vertices = voronoi(points)
/tmp/ipykernel_4300/2901581476.py:1: FutureWarning: The 'voronoi' function is considered private and will be removed in a future release.
regions, vertices = voronoi(points)
regions_df, points_df = voronoi_frames(points)
/tmp/ipykernel_4300/980072883.py:1: FutureWarning: The 'as_gdf' parameter currently defaults to True but will default to False in a future release. Set it explicitly to avoid this warning.
regions_df, points_df = voronoi_frames(points)
/tmp/ipykernel_4300/980072883.py:1: FutureWarning: The 'return_input' parameter currently defaults to True but will default to False in a future release. Set it explicitly to avoid this warning.
regions_df, points_df = voronoi_frames(points)
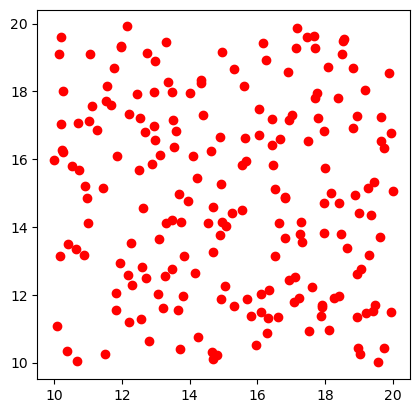
fig, ax = plt.subplots()
points_df.plot(ax=ax, color="red")
<Axes: >
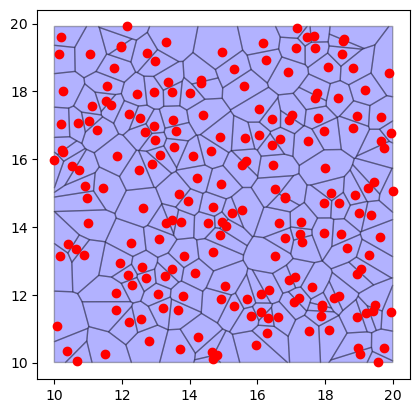
fig, ax = plt.subplots()
regions_df.plot(ax=ax, color="blue", edgecolor="black", alpha=0.3)
points_df.plot(ax=ax, color="red")
<Axes: >
Trimming¶
points = np.array(points)
maxs = points.max(axis=0)
mins = points.min(axis=0)
xr = maxs[0] - mins[0]
yr = maxs[1] - mins[1]
buff = 0.05
r = max(yr, xr) * buff
minx = mins[0] - r
miny = mins[1] - r
maxx = maxs[0] + r
maxy = maxs[1] + r
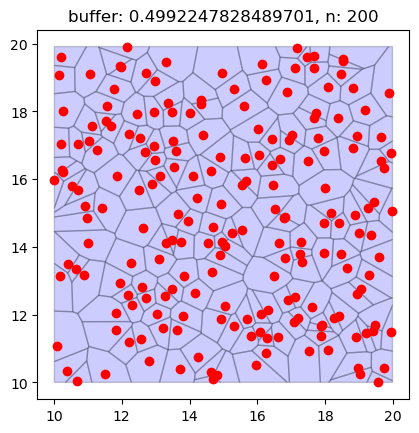
fig, ax = plt.subplots()
regions_df.plot(ax=ax, edgecolor="black", facecolor="blue", alpha=0.2)
points_df.plot(ax=ax, color="red")
plt.xlim(minx, maxx)
plt.ylim(miny, maxy)
plt.title(f"buffer: {r}, n: {n_points}")
plt.show()
Voronoi Weights¶
from libpysal.weights.contiguity import Voronoi as Vornoi_weights
w = Vornoi_weights(points)
/home/runner/micromamba/envs/test/lib/python3.14/site-packages/libpysal/weights/contiguity.py:659: FutureWarning: `use_index` defaults to False but will default to True in future. Set True/False directly to control this behavior and silence this warning
return cls.from_dataframe(region_df, **kwargs)
w.n
200
w.pct_nonzero
2.685
w.histogram
[(np.int64(1), np.int64(1)),
(np.int64(2), np.int64(6)),
(np.int64(3), np.int64(17)),
(np.int64(4), np.int64(34)),
(np.int64(5), np.int64(41)),
(np.int64(6), np.int64(63)),
(np.int64(7), np.int64(24)),
(np.int64(8), np.int64(7)),
(np.int64(9), np.int64(5)),
(np.int64(10), np.int64(1)),
(np.int64(11), np.int64(0)),
(np.int64(12), np.int64(1))]
idx = [i for i in range(w.n) if w.cardinalities[i] == 12]
points[idx]
array([[16.50851787, 13.12932895]])